Abstract
Let-7 and Lin28 establish a double-negative feedback loop to affect several biological processes, such as differentiation of stem cell, invasion and metastasis and tumorigenesis. In this study, we systematically investigated the associations between 6 potentially functional SNPs of let7 and Lin28 genes and the risk of oral cavity cancer with a case-control study including 384 oral cavity cancer cases and 731 controls. We found that the variant allele (T) of rs221636 of Lin28B was significantly associated with a reduced risk of oral cavity cancer [odds ratio (OR) = 0.73, 95% confidence interval (CI) = 0.58–0.92, P = 7.55 × 10−3 in additive model]. Bioinformatics prediction indicated that rs221636 was located at the binding site of hsa-miR-548p in the 3′ UTR of Lin28B. Luciferase activity assay also showed a lower expression level for rs221636 T allele compared with A allele. These findings indicated that rs221236 located at Lin28B may contribute to the risk of oral cavity cancer through the interruption of miRNA binding.
Similar content being viewed by others
Introduction
Oral cavity cancer is a serious worldwide public health problem, with high incidence and mortality rates. Approximately 263,900 new cases and 128,000 deaths from oral cavity cancer (including lip cancer) occurred in 2008 worldwide1. Smoking and alcohol consumption have been established as the most common environmental risk factors; however, the fact that only a small portion of exposed individuals develop oral cavity cancer suggests that genetic susceptibility plays an important role in modulating the risk of oral cavity cancer2. Therefore, the identification of susceptibility biomarkers for screening the high-risk individuals is important for the prevention of oral cavity cancer in general population.
MiRNAs are an abundant class of ~22 nucleotide noncoding RNAs that post-transcriptionally regulate the expression of protein-coding genes by targeting the 3′ untranslated region of specific messenger RNAs for degradation or translational repression3. Accumulative evidence has demonstrated the critical role of miRNAs in a variety of physiological processes, such as cell growth, cell differentiation, epithelial morphogenesis and cell survival. Furthermore, the deregulation of miRNAs has been involved in the pathogenesis of human diseases including multiple cancers3. MiRNA let-7 is the first miRNA identified in humans, originally discovered in the nematode Caenorhabditis elegans4. Let-7has been widely proposed as a tumor suppressor by regulating several oncogenes, such asK-Ras, STAT3, c-Myc and HMGA25,6,7,8,9. It has been revealed that decreased let-7 expression can increase the tumorigenicity of cancer cells10. The RNA-binding protein Lin28 is a stem cell pluripotency factor that contributes to the maintenance of stem cell characteristics and the promotion of cell malignant transformation. Recently, Lin28A and its homolog, Lin28B have been found to regulate let-7 family members through maturation process and cellular differentiation11,12. Specially, Lin28 can bind to the terminal loops of pre-let-7 elements and induce terminal uridylation of let-7 precursor miRNA, thus blocking the biogenesis of let-7 miRNAs13. Lin28A and Lin28B share similar structures; however, different functions were explored in mammalian cells14,15. For example, Lin28A suppresses let-7 biogenesis at the Dicer step in cytoplasm16, but Lin28B accumulates in the nucleus and binds pre-let-7 miRNAs to block their processing by the Microprocessor14. Because let-7 directly targets 3′UTR of Lin28A and Lin28B, this let-7/Lin28 axis establishes a double-negative feedback loop. The double negative feedback loop comprising Lin28 and let-7 has been involved in several biological processes, including differentiation of stem cell, tumorigenesis, invasion, metastasis and drug resistance and relapse13,17,18. Thus, it can be speculated that slight changes in let-7/Lin28 axis, such as sequence variants, may affect the interaction of let-7 and Lin28 and result in more significant alterations by the loop.
Many studies have shown that single nucleotide polymorphisms (SNPs) related to miRNAs may either create or disturb miRNA target interactions and induce diverse functional consequences19,20,21,22,23. Thus, in this study, we performed the genotyping of several potentially functional SNPs in let-7 and Lin28 and assessed their associations with risk of oral cavity cancer in an ongoing hospital-based case-control study of 384 cases and 731 cancer-free controls in a Chinese Han population.
Results
The selected characteristics of the cases and controls are shown in Table 1. There was no significant difference in the distributions of age, sex and smoking between the cases and controls (P = 0.163, 0.179 and 0.196, respectively). As expected, more drinkers were observed in the case group compared with that in the control group (P < 0.001). Among all oral cavity cancer cases, 341 (88.8%) were presented with squamous cell carcinoma.
Primary information and genotyping results of the 6 selected SNPs were showed in Table 2. The observed genotype frequencies of these variants were in agreement with the Hardy–Weinberg equilibrium among the controls, except rs3811463 (P = 4.94 × 10−4). The genotype frequencies of these 6 SNPs in the cases and controls are summarized in Table 3. After the adjustment for age, sex, smoking and alcohol status, multivariate logistic regression analysis revealed that rs221636 in the 3′UTR of Lin28B was significantly associated with a decreased risk of oral cavity cancer (additive model: adjusted OR = 0.73, 95% CI = 0.58–0.92, P = 7.55 × 10−3; dominant model: adjusted OR = 0.70, 95% CI = 0.53–0.91, P = 8.26 × 10−3). Even after the Bonferroni adjustment, the association remained significant (adjusted P = 0.045). Additionally, rs13293512 had a borderline association with a decreased risk of oral cavity cancer (CC vs. TT: adjusted OR = 0.68, 95% CI = 0.46–1.00, P = 4.78 × 10−2); however, the significance disappeared after the Bonferroni correction (adjusted P > 0.05). No significant association was observed between variant genotypes of the other SNPs and oral cavity cancer risk.
We further conducted the stratification analysis on the associations between rs221636 and oral cavity cancer by age, sex, smoking and drinking. As shown in Table 4, the significant association of rs221636 with oral cavity cancer risk was found among nondrinkers (adjusted OR = 0.60; 95% CI = 0.44–0.81; P = 2.30 × 10−2 for heterogeneity test) and subjects with squamous cancer (adjusted OR = 0.68, 95% CI = 0.53–0.87; P = 4.30 × 10−2 for heterogeneity test), whereas no significant differences were found between other subgroups. We then did an interaction analysis and detected a significant multiplicative interaction between rs221636 and drinking on oral cavity cancer risk (P = 1.97 × 10−2). As shown in Table 5, compared with drinkers with AA genotype, significantly decreased risks of oral cavity cancer were observed for non-drinkers with AT or TT genotypes (AT: adjusted OR = 0.24, 95%CI = 0.16–0.38; TT: adjusted OR = 0.20, 95%CI = 0.08–0.49).
To explore the functional implication of rs221636 in the development of oral cavity cancer, we used the in silico analysis tools (SNPinfo, http://snpinfo.niehs.nih.gov; PolymiRTS Database 3.0, http://compbio.uthsc.edu)24,25 to predict the potential function of this SNP and found that rs221636 was located at the target site of hsa-miR-548p, but not the let-7. Thus, we hypothesized that rs221636 might affect the expression of Lin28B by disturbing the binding of hsa-miR-548p and then the let-7/Lin28 double-negative feedback loop. To test this hypothesis, the luciferase reporter gene assay was performed and the results showed that two alleles had different effects on the expression levels of the luciferase gene when the rs221636 locus changed from the wide A allele to the variant T allele in three cell lines (Cal27, Tca8113 and 293T) (P = 2.13 × 10−6, 1.28 × 10−4 and 1.85 × 10−6, respectively) (Fig. 1). The results suggested that variant allele of rs221636 might affect the targeting of hsa-miR-548p to 3′UTR of Lin28B in oral cancer cells.
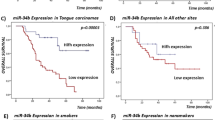
In vitro target binding assays for rs221636 A/T in Cal27, Tca8113 and 293T cell lines.
Each transfection was performed with pRL-SV40 plasmids as normalized controls. The mean fold change ± SD for plasmid with different alleles are shown after normalized by control plasmid in parallel transfection. Lin28 3′-UTR luciferase reporter plasmids (A allele or T allele) were co-transfected with chemically synthesized mature miR-548p in three cell lines. The P values are 2.13 × 10−6, 1.28 × 10−4 and 1.85 × 10−6 for Cal27, Tca8113 and 293T cell lines, respectively.
Discussion
To our knowledge, this is the first study to evaluate the effect of polymorphisms in let-7/Lin28 genes on oral cavity cancer risk in a Chinese Han population. We found that rs221636 of Lin28B, a SNP located at the binding site of some miRNA in Lin28, might affect oral cavity cancer risk through disturbing the interaction of miRNAs with Lin28.
Members of the let-7 family often promote the oncogenesis by depressing targets such as K-Ras, STAT3, c-Myc and HMGA2 in numerous types of cancer5,6,7,8,9. Specially, let-7a was down-expressed in the tissue of oral cavity cancer and might affect the metastasis and prognosis of oral cavity cancer26,27. In contrast, Lin28 and its homolog, Lin28B, are often overexpressed in primary human tumors14,15. Recent evidence has also reported that Lin28A/Lin28B block let-7 precursors from being processed to mature miRNAs, suggesting their overexpression might promote the malignancy through repression of let-728. Furthermore, studies show that let-7 represses the translation of Lin2829,30 and the knockdown of Lin28 in cell culture restores levels of mature let-7 miRNAs14,31. Thus, it is recognized that these two factors form a unique double-negative feedback, which may interact with other factors, such as RAS, MYC and NF-kB, to form a complex regulatory network and play a significant role in the tumorigenesis4,5,6,13,32,33.
Up to date, the mechanisms by which let-7/Lin28 loop homeostasis is maintained remain largely unknown. However, some studies have investigated the associations between genetic changes of let-7/Lin28 loop and the risk of human cancer. For example, Chen et al reported that a SNP rs3811463 located near the let-7 binding site in Lin28, could lead to differential regulation of Lin28 by let-7 and have a significant effect on the risk of breast cancer31. This study provided the evidence that genetic variants could directly influence the interaction of Lin28 and let-7; however, other regulatory mechanisms may also contribute to the regulation of let-7/Lin28 loop and development of cancer. In our study, we investigated the associations between several functional SNPs of let-7/Lin28 and found that another SNP of Lin28B (rs221636) might affect the risk of oral cavity cancer through disturbing the interactions of other miRNAs with Lin28, such as hsa-miR-548p. Luciferase assay also indicated that the transcription activity of reporter gene with rs221636 A allele significantly increased than that with T allele. Such results can provide more clues supporting the speculation that some genetic changes in the let-7/Lin28 loop may induce the significantly biological alterations and even the development of cancer. But, the association between rs3811463 and the risk of oral cavity cancer was non-significant in this study, which was inconsistent with the results reported in breast cancer by Chen et al., possibly because of different mechanisms involved in the development of different types of cancer. Furthermore, in this study, we also found a significant multiplicative interaction between rs221636 and drinking on oral cavity cancer risk and the decreased risk of oral cavity cancer was observed for those non-drinkers with AT or TT genotypes compared with drinkers with AA genotype. While the sample is relative small, these findings suggested that the genetic variants of Lin28B and alcohol drinking may have synergistic effect in relation to the risk of oral cavity cancer.
Some limitations are inherent in our study design. Firstly, it is a hospital-based, case-control study and inherent selection bias cannot be completely excluded. However, we applied a rigorous epidemiological design in selecting study subjects and used further statistical adjustment for known risk factors to minimize potential biases. Second, the sample size in this study (384 cases and 731 controls) is relatively small, which may have limited statistical power to detect the weak genetic effect of some SNPs. Thirdly, although we have demonstrated that rs221636 was associated with the risk of oral cavity cancer through disturbing the binding of Lin28 with other miRNAs, we are still unclear about the precise function of this SNP. Furthermore, though we have detected a significant multiplicative interaction between rs221636 variant and alcohol consumption on oral cavity cancer risk, this result is in statistical scale and future studies are required to validate this finding.
In summary, this case-control study from a Chinese population reported that the functional SNP-rs221636 of the Lin28B may modify the risk of oral cavity cancer. More rigorous studies with larger sample sizes and SNP functional relevance are warranted to replicate our findings and identify the underlying mechanism of the SNPs in the etiology of oral cavity cancer.
Methods
Ethics statement
This case-control study was approved by the institutional review board of Nanjing Medical University. Informed written consent was obtained from all subjects. The experimental protocol was carried out in accordance with the approved guidelines.
Study subjects
All newly and histologically confirmed oral cavity cancer patients were consecutively recruited from Jiangsu Stomatological Hospital and the First Affiliated Hospital of Nanjing Medical University, Nanjing, China, since January 2009 to April 2012. There were no age, sex, histology or stage restrictions, but patients with second oral cavity cancer primary tumors, primary tumors of the nasopharynx or sinonasal tract, metastasized cancer from other organs, or any histopathologic diagnosis other than oral cavity cancer were excluded. Cancer-free controls that were frequency matched to the cases on age (±5 years) and sex were randomly selected from a cohort of more than 30,000 participants in a community-based screening program for non-infectious diseases in the Jiangsu Province, China. All participants were genetically unrelated, ethnic Chinese Han population. When written informed consent was obtained, a structured questionnaire was used by trained interviewers to collect information on demographic data and environmental exposure history, such as age, sex, smoking and drinking consumption. Individuals who smoked one cigarette per day for over 1 year were considered as smokers and those who had three or more alcohol drinks a week for over 6 months were defined as alcohol drinkers. After the interview, approximately 5 ml of venous blood sample was collected from each study participant. Finally, 384 incident oral cavity cancer cases and 731 frequency-matched controls were included in this study.
SNPs selection
The dbSNP database and International HapMap Project database were first used to search all common SNPs [MAF (minor allele frequency) >0.05 in China populations] located in 3′UTR region of Lin28 genes (Lin28A and Lin28B), which is the primary binding site of miRNAs. Furthermore, the members of let-7 miRNAs (let-7a, 7b, 7c, 7d, 7e, 7f, 7g, 7i) were determined through miRBase and Gene database of NCBI. In this study, we mainly selected common SNPs located in the sequence encoding the precursors of let-7 plus 10-kb upstream region. However, no common SNP was found in the coding sequence of let-7 gene, indicating it is very conservative. Then, we used the web-based analysis tools (SNPinfo, http://snpinfo.niehs.nih.gov; PolymiRTS Database 3.0, http://compbio.uthsc.edu; TFSEARCH 1.3, http://www.cbrc.jp/research/db/TFSEARCH.html) to predict the functional implications of these SNPs24,25,34. Additionally, the linkage disequilibrium analysis was conducted to optimize the selection of SNPs (r2 > 0.8). As a result, six SNPs (Table 2) of let-7 and Lin28 were selected for genotyping.
Genotyping
Genomic DNA was extracted from a leukocyte pellet by proteinase K digestion and followed by phenol–chloroform extraction and ethanol precipitation. SNPs were genotyped by using the TaqMan allelic discrimination assay on the platform of 7900HT Real-time PCR System (Applied Biosystems, Foster City, CA). Genotyping was performed without knowing the subjects' case or control status and two negative controls (no DNA) included in each 384-well plate was used for quality control. The genotyping results were determined by using SDS 2.3 Allelic Discrimination Software (Applied Biosystems). Moreover, 10% of samples (40 cases and 70 controls) were randomly selected to repeat and the accordance rate reached 100%.
Lin28 3′-UTR promoter luciferase reporter plasmid
The Lin28 3′-UTR containing the putative recognition site rs221636 was amplified from the sample DNA, then cloned into the pMIR-REPORTTM (Applied Biosystems) vector with Mlu I and Hind III digestions. The primers were GACGCGTCACTTTGCAGGGATTA (sense) and CCAAGCTTGAGATTTCCCATGTCCTGT (antisense), which were then ligated by T4 DNA ligase (New England BioLabs) to generate the recombinant constructs. Plasmids containing the different alleles of rs221636 were generated using site-specific mutagenesis. The restriction map and sequencing were used to confirm the authenticity of all constructs in this study.
Transient transfections and luciferase assays
The Cal27, Tca8113 and 293T cells were maintained in DMEM medium supplemented with 10% heat-inactivated fetal, 10% heat-inactivated fetal bovine serum (Gibco) and 50 ug/ml streptomycin (Gibco) and incubated at 37°C in an incubator with 5% CO2. Cells were seeded at 1 × 105 cells per well in 24-well plates (BD Biosciences, Bedford, MA). Transfections were performed with cells using Lipofectamine2000 according to manufacturer's introduction (Invitrogen) after 24 h. The luciferase plasmids (empty vector for control and vectors with different rs221636 alleles) were co-transfected, respectively, into different cells with synthesized mature hsa-miR-548p mimic. The pRL-SV40 plasmid (Promega) was also co-transfected as an internal control. Six replicates for each group and the experiment repeated at least three times. After 24 hours of incubation, cells were collected and analyzed for luciferase activity with the Dual-Luciferase Reporter Assay System (Promega).
Statistical analysis
Differences in the distributions of demographic characteristics, selected variables and frequencies of the genotypes between the cases and controls were analyzed by using the χ2 test (categorical variables) and student T test (continuous variables). The associations of variant genotypes with oral cavity cancer risk were estimated by computing odds ratios (ORs) and 95% confidence intervals (CIs) from both univariate and multivariate logistic regression analyses in different genetic models. The adjustment factors for the associations included age, sex and smoking and drinking status. The Hardy-Weinberg equilibrium was tested by a goodness-offit χ2 test to compare the observed genotype frequencies with the expected ones among the control subjects. The unpaired Student's t-test was used to evaluate the associations of differences in dual-luciferase reporter gene expressions.
All the statistical analyses were performed with the Statistical Analysis System software (v.9.1.3; SAS Institute, Cary, NC). Two-sided tests were generally used for statistical analysis and P < 0.05 was considered as the level of statistical significance.
References
Jemal, A. et al. Global cancer statistics. CA Cancer J Clin 61, 69–90 (2011).
Bartsch, H., Dally, H., Popanda, O., Risch, A. & Schmezer, P. Genetic risk profiles for cancer susceptibility and therapy response. Recent Results Cancer Res 174, 19–36 (2007).
Brower, J. V., Clark, P. A., Lyon, W. & Kuo, J. S. MicroRNAs in cancer: Glioblastoma and glioblastoma cancer stem cells. Neurochem Int 77, 68–77 (2014).
Reinhart, B. J. et al. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901–906 (2000).
Sugimura, K. et al. Let-7 expression is a significant determinant of response to chemotherapy through the regulation of IL-6/STAT3 pathway in esophageal squamous cell carcinoma. Clin Cancer Res 18, 5144–5153 (2012).
Shell, S. et al. Let-7 expression defines two differentiation stages of cancer. Proc Natl Acad Sci U S A 104, 11400–11405 (2007).
Sampson, V. B. et al. MicroRNA let-7a down-regulates MYC and reverts MYC-induced growth in Burkitt lymphoma cells. Cancer Res 67, 9762–9770 (2007).
Lee, Y. S. & Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev 21, 1025–1030 (2007).
Johnson, S. M. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005).
Bussing, I., Slack, F. J. & Grosshans, H. let-7 microRNAs in development, stem cells and cancer. Trends Mol Med 14, 400–409 (2008).
Wang, Y. C. et al. Lin-28B expression promotes transformation and invasion in human hepatocellular carcinoma. Carcinogenesis 31, 1516–1522 (2010).
Viswanathan, S. R., Daley, G. Q. & Gregory, R. I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008).
Heo, I. et al. Lin28 mediates the terminal uridylation of let-7 precursor MicroRNA. Mol Cell 32, 276–284 (2008).
Piskounova, E. et al. Lin28A and Lin28B inhibit let-7 microRNA biogenesis by distinct mechanisms. Cell 147, 1066–1079 (2011).
Huang, Y. A mirror of two faces: Lin28 as a master regulator of both miRNA and mRNA. Wiley Interdiscip Rev RNA 3, 483–494 (2012).
Heo, I. et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 138, 696–708 (2009).
Esquela-Kerscher, A. & Slack, F. J. Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 6, 259–269 (2006).
Viswanathan, S. R. & Daley, G. Q. Lin28: A microRNA regulator with a macro role. Cell 140, 445–449 (2010).
Jiang, Y. et al. Evaluation of genetic variants in microRNA biosynthesis genes and risk of breast cancer in Chinese women. Int J Cancer 133, 2216–2224 (2013).
Jiang, Y. et al. Genetic variation in a hsa-let-7 binding site in RAD52 is associated with breast cancer susceptibility. Carcinogenesis 34, 689–693 (2013).
Ma, H. et al. Genetic variations in key microRNA processing genes and risk of head and neck cancer: a case-control study in Chinese population. PLoS One 7, e47544 (2012).
Vaishnavi, V., Manikandan, M. & Munirajan, A. K. Mining the 3′UTR of autism-implicated genes for SNPs perturbing microRNA regulation. Genomics Proteomics Bioinformatics 12, 92–104 (2014).
Han, S. J. et al. Kaposi's sarcoma-associated herpesvirus microRNA single-nucleotide polymorphisms identified in clinical samples can affect microRNA processing, level of expression and silencing activity. J Virol 87, 12237–12248 (2013).
Xu, Z. & Taylor, J. A. SNPinfo: integrating GWAS and candidate gene information into functional SNP selection for genetic association studies. Nucleic Acids Res 37, W600–6005 (2009).
Bhattacharya, A., Ziebarth, J. D. & Cui, Y. PolymiRTS Database 3.0: linking polymorphisms in microRNAs and their target sites with human diseases and biological pathways. Nucleic Acids Res 42, D86–91 (2014).
Yu, C. C. et al. MicroRNA let-7a represses chemoresistance and tumourigenicity in head and neck cancer via stem-like properties ablation. Oral Oncol 47, 202–210 (2011).
Chang, C. J. et al. Let-7d functions as novel regulator of epithelial-mesenchymal transition and chemoresistant property in oral cancer. Oncol Rep 26, 1003–1010 (2011).
Viswanathan, S. R. et al. Lin28 promotes transformation and is associated with advanced human malignancies. Nat Genet 41, 843–848 (2009).
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat Cell Biol 10, 987–993 (2008).
Yang, X. et al. Double-negative feedback loop between reprogramming factor LIN28 and microRNA let-7 regulates aldehyde dehydrogenase 1-positive cancer stem cells. Cancer Res 70, 9463–9472 (2010).
Chen, A. X. et al. Germline genetic variants disturbing the Let-7/LIN28 double-negative feedback loop alter breast cancer susceptibility. PLoS Genet 7, e1002259 (2011).
Dangi-Garimella, S. et al. Raf kinase inhibitory protein suppresses a metastasis signalling cascade involving LIN28 and let-7. EMBO J 28, 347–358 (2009).
Iliopoulos, D., Hirsch, H. A. & Struhl, K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA and IL6 links inflammation to cell transformation. Cell 139, 693–706 (2009).
Heinemeyer, T. et al. Databases on transcriptional regulation: TRANSFAC, TRRD and COMPEL. Nucleic Acids Res 26, 362–367 (1998).
Acknowledgements
This work was partly supported by National Natural Science Foundation of China (81302361 and 81473048), Specialized Research Fund for the Doctoral Program of Higher Education of China (20133234120013), China Postdoctoral Science Foundation (2013M540457), Jiangsu Planned Projects for Postdoctoral Research Funds (1301018A) and Priority Academic Program for the Development of Jiangsu Higher Education Institutions [PAPD, 2014-37 and Public Health and Preventive Medicine].
Author information
Authors and Affiliations
Contributions
H.Y., H.M. and N.C. conceived and designed the experiments. Y.Z., L.Z. and R.W. performed the experiment. H.Y., R.W. and H.J. analyzed the data. R.W., L.Z. and L.M. prepared the samples. Y.Z., R.W. and L.Z. wrote the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Zhang, Y., zhu, L., Wang, R. et al. Genetic variants in let-7/Lin28 modulate the risk of oral cavity cancer in a Chinese Han Population. Sci Rep 4, 7434 (2014). https://doi.org/10.1038/srep07434
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07434
This article is cited by
-
The pluripotency factor LIN28B is involved in oral carcinogenesis and associates with tumor aggressiveness and unfavorable prognosis
Cancer Cell International (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.