Abstract
Background
The expression of Fos-related antigen 1 (Fra-1) affects tumor progression, migration, and invasion. In this study, we identified the genes regulated by Fra-1 in esophageal squamous cell carcinoma (ESCC).
Methods
We constructed Fra-1 knockdown models via the transfection of small interfering RNA (siRNA) into ESCC cell lines (TE10, TE11). The expression levels of the genes in the knockdown models were analyzed using a microarray and a Biobase Upstream Analysis, while the expression levels of the candidate genes in the primary tumors of surgical specimens obtained from ESCC patients were determined using real-time polymerase chain reaction (PCR) and immunohistochemical staining. The clinicopathological features were then analyzed.
Results
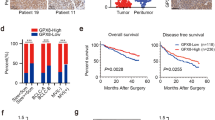
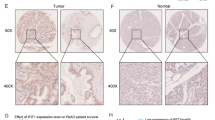
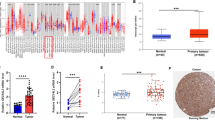
The Biobase Upstream Analysis showed the high-mobility-group protein-1 (HMGA1) to be a significant gene regulated by Fra-1. Actual binding of Fra-1 to the promotor region of HMGA1 was revealed in subsequent chromatin immunoprecipitation PCR experiments. Patients with a positive HMGA1 expression had a poor prognosis, and a multivariate analysis demonstrated a positive HMGA1 expression to be a significant independent prognostic factor.
Conclusion
HMGA1 is regulated by Fra-1 in ESCC, and the HMGA1 expression is significantly associated with a poor prognosis in ESCC patients. Downregulation of the HMGA1 expression may become a practical treatment strategy against ESCC in the future.




Similar content being viewed by others
References
Courrech Staal EF, van Coevorden F, Cats A, et al. Outcome of low-volume surgery for esophageal cancer in a high-volume referral center. Ann Surg Oncol. 2009;16:3219–26.
Law SY, Fok M, Chen SW. Wong J. A comparison of outcome after resection for sequamous cell carcinomas and adenocarcinomas of the esophagus and cardia. Surg Gynecol Obstet. 1992;175:107–12.
Curran T, Franza BR Jr. Fos and Jun: the AP-1 connection. Cell. 1998;55:395–7.
Angel P, Karin M. Specific members of the Jun protein family regulate collagenase expression in response to various extracellular stimuli. Matrix Suppl. 1992;1:156–4.
Mechta F, Lallemand D, Pfarr CM, et al. Transformation by ras modifies AP1 composition and activity. Oncogene. 1997;14:837–7.
Kakumoto K, Sasai K, Sukezawa T, et al. FRA1 is a determinant for the difference in RAS-induced transformation between human and rat fibroblasts. Proc Natl Acad Sci USA. 2006;103:5490–5.
Usui A, Hoshino I, Akutsu Y, et al. The molecular role of Fra-1 and its prognostic significance in human esophageal squamous cell carcinoma. Cancer. 2012;118:3387–6.
Akanuma N, Hoshino I, Akutsu Y, et al. MicroRNA-133a regulates the mRNAs of two invadopodia-related proteins, FSCN1 and MMP14, in esophageal cancer. Br J Cancer. 2014;110:189–8.
Nishihira T, Hashimoto Y, Katayama M, et al. Molecular and cellular features of esophageal cancer cells. J Cancer Res Clin Oncol. 1993;119:441–9.
Kel AE, Gobling E, Reuter I, et al. MATCH: a tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Res. 2003;31:3576–9.
Wingender E, Chen X, Hehl R, et al. TRANSFAC: an intergrade system for gene expression regulation. Nucleic Acids Res. 2000;28:316–9.
Ameyar-Zazoua M, Wisniewska MB, Bakiri L, et al. AP-1 dimers regulate transcription of the p14/p19ARF tumor suppressor gene. Oncogene. 2005;24:2298–6.
Belfuise K, Kersual N, Galtier F, et al. FRA-1 expression level regulates proliferation and invasiveness of breast cancer cells. Oncogene. 2005;24:1434–4.
Burch PM, Yuan Z, Loonen A, et al. An extracellular sgnal-regulated kinase1- and 2-dependent program of chromatin trafficking of c-Fos and Fra-1 is required for cyclin D1 expression during cell cycle reentry. Mol Cell Biol. 2004;24:4696–9.
Zhang Q, Adiseshaiah P, Reddv SP. Matrix metalloproteinase/epidermal growth factor receptor/mitogen-activated protein kinase signaling regulate fra-1 induction by cigarette smoke in lung epithelial cells. Am J Respir Cell Mol Biol. 2005;32:72–1.
Hommura F, Katabami M, Leaner VD, et al. HMG-I/Y is a c-Jun/activator protein-1 target gene and is necessary for c-Jun-induced anchorage-independent growth in Rat1a cells. Mol Cancer Res. 2004;2:305–4.
Wood LJ, Mukherjee M, Dolde CE, et al. HMG-I/Y: a new c-Myc target gene and potential human oncogene. Mol Cell Biol. 2000;20:5490–2.
Dhar A, Hu J, Reeves R, et al. Dominant negative c-Jun (TAM67) target genes: HMGA1 is required for tumor promoter-induced transformation. Oncogene. 2004;23:4466–6.
Giannini G, Cerignoli F, Mellone M, et al. High mobility group A1 is a molecular target for MYCN in human neuroblastoma. Cancer Res. 2005;65:8308–6.
Giannini G, Cerignoli F, Mellone M, et al. Molecular mechanism of HMGA1 deregulation in human neuroblastoma. Cancer Lett. 2005;228:97–104.
Pedulla ML, Treff NR, Resar LM, et al. Sequence and analysis of the murine Hmgiy (Hmga1) gene locus. Gene. 2001;271:51–8.
Sgarra R, Rustighi A, Tessari MA, et al. Nuclear phosphoproteins HMGA and their relationship with chromatin structure and cancer. FEBS Lett. 2004;574:1–8.
Johnson KR, Cook SA, Davisson MT. Chromosomal localization of the murine gene and two related sequences encoding high-mobility-group I and Y proteins. Genomics. 1992;12:503–9.
Mather JF, Nathans D. Mulrivalent DNA-binding properties of the HMG-1 proteins. Proc Natl Acad Sci USA. 1996;93:6716–20.
Grosschedl R, Giese K, Pagel J. HMG domain proteins: architectural elements in the assembly of nucleoprotein structures. Trends Genet. 1994;10:94–100.
Chiappetta G, Avantaggiato V, Visconti R, et al. High level expression of the HMGI (Y) gene during embryonic development. Oncogene. 1996;13:2439–6.
Czyz W, Balcerczak E, Jakubiak M, et al. HMGI(Y) gene expression as a potential marker of thyroid follicular carcinoma. Langenbecks Arch Surg. 2004;389:193–7.
Tamimi Y, van der Poel HG, Denyn MM, et al. Increased expression of high mobility group protein I(Y) in high grade prostatic cancer determined by in situ hybridization. Cancer Res. 1993;53:5512–6.
Fedele M, Bandiera A, Chiappetta G, et al. Human colorectal carcinomas express high levels of high mobility group HMGI(Y) proteins. Cancer Res. 1996;56:1896–901.
Nam ES, Kim DH, Cho SJ, et al. Expression of HMGI(Y) associated with malignant phenotype of human gastric tissue. Histopathology. 2003;42:466–71.
Abe N, Watanabe T, Masaki T, et al. Pancreatic duct cell carcinomas express high levels of high mobility group I(Y) proteins. Cancer Res. 2000;60:3117–22.
Rho YS, Lim YC, Park IS, et al. High mobility group HMGI(Y) protein expression in head and neck squamous cell carcinoma. Acta Otolaryngol. 2007;127:76–81.
Chen X, Lechago J, Ertan A, et al. Expression of the high mobility group protein HMGI(Y) correlates with malignant progression in Barrett’s metaplasia. Cancer Epidemiol Biomarkers Prev. 2004;13:30–3.
Di Cello F, Shin J, Harborm K, et al. Knockdown of HMGA1 inhibits human breast cancer cell growth and metastasis in immunodeficient mice. Biochem Biophys Res Commun. 2013;434:70–4.
Dolde CE, Mukherjee M, Cho C, et al. HMG-I/Y in human breast cancer cell lines. Breast Cancer Res Treat. 2002;71:181–91.
Belton A, Gabrovsky A, Bae YK, et al. HMGA1 induces intestinal polyposis in transgenic mice and drives tumor progression and stem cell properties in colon cancer cells. PLoS One. 2012;7:e30034.
Funding
The authors received no funding support for this study.
Disclosure
Takeshi Toyozumi, Isamu Hoshino, Masahiko Takahashi, Akihiro Usui, Yasunori Akutsu, Naoyuki Hanari, Kentaro Murakami, Masayuki Kano, Naoki Akanuma, Hiroshi Suitoh, Yasunori Matsumoto, Nobuhumi Sekino, Aki Komatsu, and Hisahiro Matsubara have no disclosures to declare.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Toyozumi, T., Hoshino, I., Takahashi, M. et al. Fra-1 Regulates the Expression of HMGA1, Which is Associated with a Poor Prognosis in Human Esophageal Squamous Cell Carcinoma. Ann Surg Oncol 24, 3446–3455 (2017). https://doi.org/10.1245/s10434-016-5666-5
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-016-5666-5