Abstract
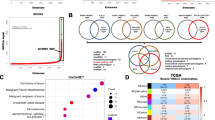
Objective: In this study, we aimed to expand current knowledge of head and neck squamous cell carcinoma (HNSCC)-associated long noncoding RNAs (lncRNAs), and to discover potential lncRNA prognostic biomarkers for HNSCC based on next-generation RNA-seq. Methods: RNA-seq data of 546 samples from patients with HNSCC were downloaded from The Cancer Genome Atlas (TCGA), including 43 paired samples of tumor tissue and adjacent normal tissue. An integrated analysis incorporating differential expression, weighted gene co-expression networks, functional enrichment, clinical parameters, and survival analysis was conducted to discover HNSCC-associated lncRNAs. The function of CYTOR was verified by cell-based experiments. To further identify lncRNAs with prognostic significance, a multivariate Cox proportional hazard regression analysis was performed. The identified lncRNAs were validated with an independent cohort using clinical feature relevance analysis and multivariate Cox regression analysis. Results: We identified nine HNSCC-relevant lncRNAs likely to play pivotal roles in HNSCC onset and development. By functional enrichment analysis, we revealed that CYTOR might participate in the multistep pathological processes of cancer, such as ribosome biogenesis and maintenance of genomic stability. CYTOR was identified to be positively correlated with lymph node metastasis, and significantly negatively correlated with overall survival (OS) and disease free survival (DFS) of HNSCC patients. Moreover, CYTOR inhibited cell apoptosis following treatment with the chemotherapeutic drug diamminedichloroplatinum (DDP). HCG22, the most dramatically down-regulated lncRNA in tumor tissue, may function in epidermis differentiation. It was also significantly associated with several clinical features of patients with HNSCC, and positively correlated with patient survival. CYTOR and HCG22 maintained their prognostic values independent of several clinical features in multivariate Cox hazards analysis. Notably, validation either based on an independent HNSCC cohort or by laboratory experiments confirmed these findings. Conclusions: Our transcriptomic analysis suggested that dysregulation of these HNSCC-associated lncRNAs might be involved in HNSCC oncogenesis and progression. Moreover, CYTOR and HCG22 were confirmed as two independent prognostic factors for HNSCC patient survival, providing new insights into the roles of these lncRNAs in HNSCC as well as clinical applications.
摘要
目的
研究长链非编码RNA(lncRNA)与头颈部肿瘤 发生、发展及预后的关系。
创新点
通过使用整合的转录组分析方法筛选出与头颈部 肿瘤密切相关的lncRNA,其中CYTOR和HCG22 在头颈部肿瘤发生发展中具有重要的生物学功 能和临床预后价值,为制定新的治疗策略和探索 新的预后标记分子提供参考。
方法
从癌症基因组数据集(The Cancer Genome Atlas) 中获得RNA-seq 数据。结合差异表达分析和共 表达网络分析的方法发掘出与头颈部鳞状细胞 癌相关的lncRNA,探讨其与头颈部肿瘤临床病 理变化和预后的关系,进一步利用外部数据集以 及细胞水平进行验证。
结论
发现9 个与头颈部肿瘤发生发展密切相关的 lncRNA,其中CYTOR 可能参与核糖体的生物合 成,与病人生存率呈负相关。HCG22 可能参与 细胞表皮分化过程,与病人生存率呈正相关。此 外,CYTOR 和HCG22 可作为头颈部鳞状细胞癌 独立的预后标记物。
Similar content being viewed by others
References
Aken BL, Ayling S, Barrell D, et al., 2016. The Ensembl gene annotation system. Database, 2016:baw093. https://doi.org/10.1093/database/baw093
Anders S, Pyl PT, Huber W, 2015. HTSeq—a python framework to work with high–throughput sequencing data. Bioinformatics, 31(2):166–169. https://doi.org/10.1093/bioinformatics/btu638
Barna M, Pusic A, Zollo O, et al., 2008. Suppression of Myc oncogenic activity by ribosomal protein haploinsufficiency. Nature, 456(7224):971–975. https://doi.org/10.1038/nature07449
Bartkova J, Hořejší Z, Koed K, et al., 2005. DNA damage response as a candidate anti–cancer barrier in early human tumorigenesis. Nature, 434(7035):864–870. https://doi.org/10.1038/nature03482
Bartonicek N, Maag JLV, Dinger ME, 2016. Long noncoding RNAs in cancer:mechanisms of action and technological advancements. Mol Cancer, 15(1):43. https://doi.org/10.1186/s12943-016-0530-6
Benjamini Y, Hochberg Y, 1995. Controlling the false discovery rate:a practical and powerful approach to multiple testing. J Roy Statist Soc B, 57(1):289–300.
Bhatt AN, Mathur R, Farooque A, et al., 2010. Cancer biomarkers—current perspectives. Indian J Med Res, 132:129–149.
Cheetham SW, Gruhl F, Mattick JS, et al., 2013. Long noncoding RNAs and the genetics of cancer. Br J Cancer, 108(12):2419–2425. https://doi.org/10.1038/bjc.2013.233
Chen WM, Huang MD, Sun DP, et al., 2016. Long intergenic non–coding RNA 00152 promotes tumor cell cycle progression by binding to EZH2 and repressing p15 and p21 in gastric cancer. Oncotarget, 7(9):9773–9787.https://doi.org/10.18632/oncotarget.6949
Chi LM, Lee CW, Chang KP, et al., 2009. Enhanced interferon signaling pathway in oral cancer revealed by quantitative proteome analysis of microdissected specimens using 16O/18O labeling and integrated two–dimensional LC–ESIMALDI tandem MS. Mol Cell Proteomics, 8(7):1453–1474. https://doi.org/10.1074/mcp.M800460-MCP200
de Lena PG, Paz–Gallardo A, Paramio JM, et al., 2017. Clusterization in head and neck squamous carcinomas based on lncRNA expression:molecular and clinical correlates. Clin Epigenetics, 9:36. https://doi.org/10.1186/s13148-017-0334-6
Derrien T, Johnson R, Bussotti G, et al., 2012. The GENCODE v7 catalog of human long noncoding RNAs:analysis of their gene structure, evolution, and expression. Genome Res, 22(9):1775–1789. https://doi.org/10.1101/gr.132159.111
Eales KL, Hollinshead KER, Tennant DA, 2016. Hypoxia and metabolic adaptation of cancer cells. Oncogenesis, 5(1):e190. https://doi.org/10.1038/oncsis.2015.50
Engreitz JM, Ollikainen N, Guttman M, 2016. Long noncoding RNAs:spatial amplifiers that control nuclear structure and gene expression. Nat Rev Mol Cell Biol, 17(12):756–770. https://doi.org/10.1038/nrm.2016.126
Feng L, Houck JR, Lohavanichbutr P, et al., 2017. Transcriptome analysis reveals differentially expressed lncRNAs between oral squamous cell carcinoma and healthy oral mucosa. Oncotarget, 8(19):31521–31531. https://doi.org/10.18632/oncotarget.16358
Gabay M, Li YL, Felsher DW, 2014. MYC activation is a hallmark of cancer initiation and maintenance. Cold Spring Harb Perspect Med, 4(6):a014241. https://doi.org/10.1101/cshperspect.a014241
Geng YJ, Xie SL, Li Q, et al., 2011. Large intervening non–coding RNA HOTAIR is associated with hepatocellular carcinoma progression. J Int Med Res, 39(6):2119–2128. https://doi.org/10.1177/147323001103900608
Gold KA, Lee HY, Kim ES, 2009. Targeted therapies in squamous cell carcinoma of the head and neck. Cancer, 115(5):922–935. https://doi.org/10.1002/cncr.24123
Hajjari M, Salavaty A, 2015. HOTAIR:an oncogenic long non–coding RNA in different cancers. Cancer Biol Med, 12(1):1–9. https://doi.org/10.7497/j.issn.2095-3941.2015.0006
Harrow J, Frankish A, Gonzalez JM, et al., 2012. GENCODE:the reference human genome annotation for the encode project. Genome Res, 22(9):1760–1774. https://doi.org/10.1101/gr.135350.111
Holoch D, Moazed D, 2015. RNA–mediated epigenetic regulation of gene expression. Nat Rev Genet, 16(2):71–84. https://doi.org/10.1038/nrg3863
Huarte M, 2015. The emerging role of lncRNAs in cancer. Nat Med, 21(11):1253–1261. https://doi.org/10.1038/nm.3981
Kassambara A, Kosinski M, 2017. Survminer:Drawing Survival Curves Using ‘ggplot2’. R Package Version 0.4.0. https://CRAN.R-project.org/package=survminer [Accessed on June 10, 2017].
Kim K, Jutooru I, Chadalapaka G, et al., 2013. HOTAIR is a negative prognostic factor and exhibits pro–oncogenic activity in pancreatic cancer. Oncogene, 32(13):1616–1625. https://doi.org/10.1038/onc.2012.193
Kinsella RJ, Kähäri A, Haider S, et al., 2011. Ensembl BioMarts:a hub for data retrieval across taxonomic space. Database, 2011:bar030. https://doi.org/10.1093/database/bar030
Kohl M, Wiese S, Warscheid B, 2011. Cytoscape:software for visualization and analysis of biological networks. In: Hamacher M, Eisenacher M, Stephan C (Eds.), Data Mining in Proteomics. Humana Press, p.291–303. https://doi.org/10.1007/978-1-60761-987-1_18
Lamouille S, Xu J, Derynck R, 2014. Molecular mechanisms of epithelial–mesenchymal transition. Nat Rev Mol Cell Biol, 15(3):178–196. https://doi.org/10.1038/nrm3758
Langfelder P, Horvath S, 2008. WGCNA:an R package for weighted correlation network analysis. BMC Bioinform, 9:559. https://doi.org/10.1186/1471-2105-9-55.
Law CW, Chen Y, Shi W, et al., 2014. voom:precision weights unlock linear model analysis tools for RNA–seq read counts. Genome Biol, 15(2):R29. https://doi.org/10.1186/gb-2014-15-2-r29
LeBleu VS, O'Connell JT, Gonzalez Herrera KN, et al., 2014. PGC–1α mediates mitochondrial biogenesis and oxidative phosphorylation in cancer cells to promote metastasis. Nat Cell Biol, 16(10):992–1003. https://doi.org/10.1038/ncb3039
Leemans CR, Braakhuis BJM, Brakenhoff RH, 2011. The molecular biology of head and neck cancer. Nat Rev Cancer, 11(1):9–22. https://doi.org/10.1038/nrc2982
Li DD, Feng JP, Wu TY, et al., 2013. Long intergenic noncoding RNA HOTAIR is overexpressed and regulates PTEN methylation in laryngeal squamous cell carcinoma. Am J Pathol, 182(1):64–70. https://doi.org/10.1016/j.ajpath.2012.08.042
Li X, Wu Z, Mei Q, et al., 2013. Long non–coding RNA HOTAIR, a driver of malignancy, predicts negative prognosis and exhibits oncogenic activity in oesophageal squamous cell carcinoma. Br J Cancer, 109(8):2266–2278. https://doi.org/10.1038/bjc.2013.548
Liu BD, Sun LJ, Liu Q, et al., 2015. A cytoplasmic NF–κB interacting long noncoding RNA blocks IκB phosphorylation and suppresses breast cancer metastasis. Cancer Cell, 27(3):370–381. https://doi.org/10.1016/j.ccell.2015.02.004
Liu R, Cheng Y, Yu J, et al., 2015. Identification and validation of gene module associated with lung cancer through coexpression network analysis. Gene, 563(1):56–62. https://doi.org/10.1016/j.gene.2015.03.008
Mäbert K, Cojoc M, Peitzsch C, et al., 2014. Cancer biomarker discovery:current status and future perspectives. Int J Radiat Biol, 90(8):659–677. https://doi.org/10.3109/09553002.2014.892229
McCarthy DJ, Chen YS, Smyth GK, 2012. Differential expression analysis of multifactor RNA–Seq experiments with respect to biological variation. Nucleic Acids Res, 40(10):4288–4297. https://doi.org/10.1093/nar/gks042
Miller DL, Davis JW, Taylor KH, et al., 2015. Identification of a human papillomavirus–associated oncogenic miRNA panel in human oropharyngeal squamous cell carcinoma validated by bioinformatics analysis of the cancer genome atlas. Am J Pathol, 185(3):679–692. https://doi.org/10.1016/j.ajpath.2014.11.018
Min SN, Wei T, Wang XT, et al., 2017. Clinicopathological and prognostic significance of homeobox transcript antisense RNA expression in various cancers:a meta–analysis. Medicine (Baltimore), 96(23):e7084. https://doi.org/10.1097/MD.0000000000007084
Nohata N, Abba MC, Gutkind JS, 2016. Unraveling the oral cancer lncRNAome:identification of novel lncRNAs associated with malignant progression and HPV infection. Oral Oncol, 59:58–66. https://doi.org/10.1016/j.oraloncology.2016.05.014
Nötzold L, Frank L, Gandhi M, et al., 2017. The long non–coding RNA LINC00152 is essential for cell cycle progression through mitosis in HeLa cells. Sci Rep, 7:2265. https://doi.org/10.1038/s41598-017-02357-0
Parshall MB, 2013. Unpacking the 2×2 table. Hear Lung J Acute Crit Care, 42(3):221–226. https://doi.org/10.1016/j.hrtlng.2013.01.006
Peng WX, Koirala P, Mo YY, 2017. LncRNA–mediated regulation of cell signaling in cancer. Oncogene, 36(41):5661–5667. https://doi.org/10.1038/onc.2017.184
Posner MR, Hershock DM, Blajman CR, et al., 2007. Cisplatin and fluorouracil alone or with docetaxel in head and neck cancer. New Engl J Med, 357(17):1705–1715. https://doi.org/10.1056/nejmoa070956
Pritzker KPH, 2015. Predictive and prognostic cancer biomarkers revisited. Expert Rev Mol Diagn, 15(8):971–974. https://doi.org/10.1586/14737159.2015.1063421
Quek XC, Thomson DW, Maag JLV, et al., 2015. LncRNAdb v2.0:expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res, 43(D1):D168–D173. https://doi.org/10.1093/nar/gku988
R Development Core Team, 2011. R:A Language and Environment for Statistical Computing. R Development Core Team, Vienna, Austria.
Rhodes DR, Yu JJ, Shanker K, et al., 2004. ONCOMINE:a cancer microarray database and integrated data–mining platform. Neoplasia, 6(1):1–6. https://doi.org/10.1016/S1476-5586(04)80047-2
Rickman DS, Millon R, de Reynies A, et al., 2008. Prediction of future metastasis and molecular characterization of head and neck squamous–cell carcinoma based on transcriptome and genome analysis by microarrays. Oncogene, 27(51):6607–6622. https://doi.org/10.1038/onc.2008.251
Robinson MD, McCarthy DJ, Smyth GK, 2010. edgeR:a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics, 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Ruggero D, Pandolfi PP, 2003. Does the ribosome translate cancer? Nat Rev Cancer, 3(3):179–192. https://doi.org/10.1038/nrc1015
Salazar C, Calvopiña D, Punyadeera C, 2014. miRNAs in human papilloma virus associated oral and oropharyngeal squamous cell carcinomas. Expert Rev Mol Diagn, 14(8):1033–1040. https://doi.org/10.1586/14737159.2014.960519
Salyakina D, Tsinoremas NF, 2016. Non–coding RNAs profiling in head and neck cancers. NPJ Genomic Med, 1:15004. https://doi.org/10.1038/npjgenmed.2015.4
Schmitt AM, Chang HY, 2016. Long noncoding RNAs in cancer pathways. Cancer Cell, 29(4):452–463. https://doi.org/10.1016/j.ccell.2016.03.010
Schmitt AM, Garcia JT, Hung T, et al., 2016. An inducible long noncoding RNA amplifies DNA damage signaling. Nat Genet, 48(11):1370–1376. https://doi.org/10.1038/ng.3673
Seiwert TY, Salama JK, Vokes EE, 2007. The chemoradiation paradigm in head and neck cancer. Nat Clin Pract Oncol, 4(3):156–171. https://doi.org/10.1038/ncponc0750
Signal B, Gloss BS, Dinger ME, 2016. Computational approaches for functional prediction and characterisation of long noncoding RNAs. Trends Genet, 32(10):620–637. https://doi.org/10.1016/j.tig.2016.08.004
Song L, Langfelder P, Horvath S, 2012. Comparison of co–expression measures:mutual information, correlation, and model based indices. BMC Bioinformatics, 13:328. https://doi.org/10.1186/1471-2105-13-32.
Subramanian A, Tamayo P, Mootha VK, et al., 2005. Gene set enrichment analysis:a knowledge–based approach for interpreting genome–wide expression profiles. Proc Natl Acad Sci USA, 102(43):15545–15550. https://doi.org/10.1073/pnas.0506580102
Supek F, Bošnjak M, Škunca N, et al., 2011. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE, 6(7):e21800. https://doi.org/10.1371/journal.pone.0021800
Therneau T, 2017. A Package for Survival Analysis in S. R Package Version 2.41–2. https://CRAN.R-project.org/package=survival [Accessed on Mar. 20, 2017].
Tsai MC, Manor O, Wan Y, et al., 2010. Long noncoding RNA as modular scaffold of histone modification complexes. Science, 329(5992):689–693. https://doi.org/10.1126/science.1192002
van Riggelen J, Yetil A, Felsher DW, 2010. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat Rev Cancer, 10(4):301–309. https://doi.org/10.1038/nrc2819
Wade M, Wahl GM, 2006. c–Myc, genome instability, and tumorigenesis:the devil is in the details. Curr Top Microbiol Immunol, 302:169–203. https://doi.org/10.1007/3-540-32952-8_7
Wilson WR, Hay MP, 2011. Targeting hypoxia in cancer therapy. Nat Rev Cancer, 11(6):393–410. https://doi.org/10.1038/nrc3064
Yan L, Zhan C, Wu JH, et al., 2016. Expression profile analysis of head and neck squamous cell carcinomas using data from The Cancer Genome Atlas. Mol Med Rep, 13(5):4259–4265. https://doi.org/10.3892/mmr.2016.5054
Yates A, Akanni W, Amode MR, et al., 2016. Ensembl 2016. Nucleic Acids Res, 44(D1):D710–D716. https://doi.org/10.1093/nar/gkv1157
Yu GC, Wang LG, Han YY, et al., 2012. clusterProfiler:an R package for comparing biological themes among gene clusters. OMICS A J Integr Biol, 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Yu JJ, Liu Y, Guo C, et al., 2017. Upregulated long non–coding RNA LINC00152 expression is associated with progression and poor prognosis of tongue squamous cell carcinoma. J Cancer, 8(4):523–530. https://doi.org/10.7150/jca.17510
Zaidi MR, Davis S, Noonan FP, et al., 2011. Interferon–γ links ultraviolet radiation to melanomagenesis in mice. Nature, 469(7331):548–553. https://doi.org/10.1038/nature09666
Zhang B, Horvath S, 2005. A general framework for weighted gene co–expression network analysis. Stat Appl Genet Mol Biol, 4(1):Article 17. https://doi.org/10.2202/1544-6115.1128
Zhang SC, Tian LL, Ma PH, et al., 2015. Potential role of differentially expressed lncRNAs in the pathogenesis of oral squamous cell carcinoma. Arch Oral Biol, 60(10):1581–1587. https://doi.org/10.1016/j.archoralbio.2015.08.003
Zhao J, Liu YC, Zhang WH, et al., 2015. Long non–coding RNA Linc00152 is involved in cell cycle arrest, apoptosis, epithelial to mesenchymal transition, cell migration and invasion in gastric cancer. Cell Cycle, 14(19):3112–3123. https://doi.org/10.1080/15384101.2015.1078034
Author information
Authors and Affiliations
Corresponding author
Additional information
Project supported by the National Natural Science Foundation of China (Nos. 31471226 and 91440108) and the Fundamental Research Funds for the Central Universities (Nos. WK2070000044 and WK2070000034), China
Electronic supplementary materials: The online version of this article (https://doi.org/10.1631/jzus.B1700319) contains supplementary materials, which are available to authorized users
Electronic supplementary material
11585_2018_300_MOESM1_ESM.pdf
Transcriptomic analysis reveals key lncRNAs associated with ribosomal biogenesis and epidermis differentiation in head and neck squamous cell carcinoma
Rights and permissions
About this article
Cite this article
Guo, Yz., Sun, Hh., Wang, Xt. et al. Transcriptomic analysis reveals key lncRNAs associated with ribosomal biogenesis and epidermis differentiation in head and neck squamous cell carcinoma. J. Zhejiang Univ. Sci. B 19, 674–688 (2018). https://doi.org/10.1631/jzus.B1700319
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1631/jzus.B1700319
Key words
- Head and neck squamous cell carcinoma
- Long noncoding RNA (lncRNA)
- Weighted gene co-expression network analysis (WGCNA)
- Clinicopathological feature
- Multivariate Cox regression model